details
Our work is currently focused on four topics:
- Protein transport and protein assembly mechanisms in bacterial cells.
Bacteria like eukaryotic cells have to export a large subset of the cytosolically synthesized proteins into the inner membrane, the periplasmic space, the outer membrane or the extracellular space. To achieve this, bacteria have developed highly specific and highly sophisticated protein transport machineries. Some of these pathways are found in bacteria only and are crucial determinants for bacterial pathogenicity; other pathways were originally “invented” by prokaryotes but have been conserved throughout evolution and are present also in higher eukaryotes. Therefore bacteria are perfect model organisms for studying the molecular details of these essential processes.
Our work on protein transport and assembly is currently funded by the German Science Foundation (DFG) and within the Priority Program SPP 2002 (http://www.spp2002.uni-kiel.de/wordpress/)
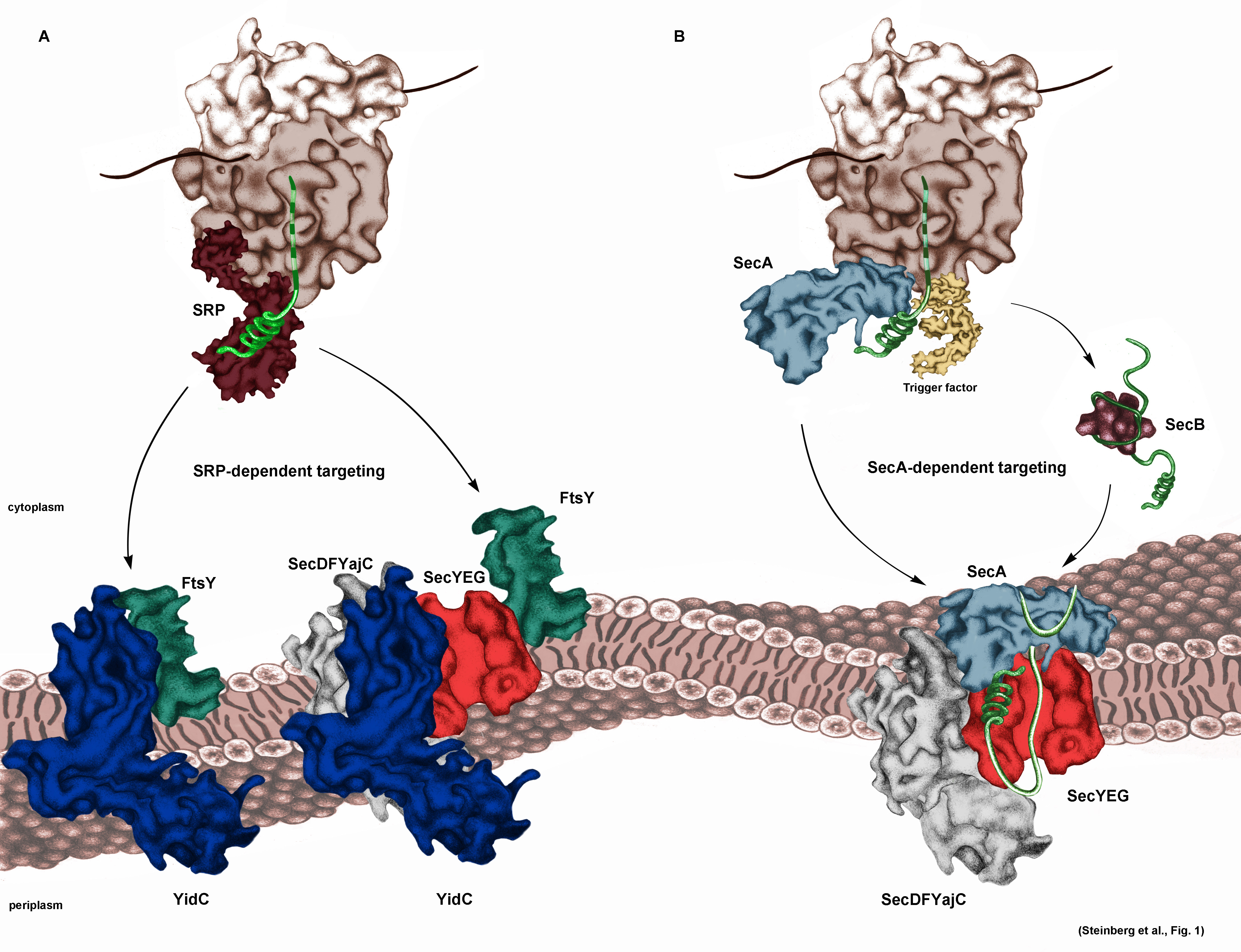
Currently studied protein transport systems in bacterial cells
- Alternative mechanisms of protein localization.
The classical concept of protein transport is based on the idea that the targeting information is retained within the protein itself; this is summarized in the so called Signal Sequence Hypothesis, which earned Gunter Blobel the Nobelprize; https://www.nobelprize.org/prizes/medicine/1999/summary/.
However, alternatively, the targeting information can also be retained within the mRNA. While multiple examples for specific mRNA localization exist in eukaryotic cells, in bacteria this is still largely a terra incognita; which is in part related to the small size of bacterial cells. We are currently testing a single-molecule set-up for studying localization and movements of single mRNA molecules in bacterial cells:
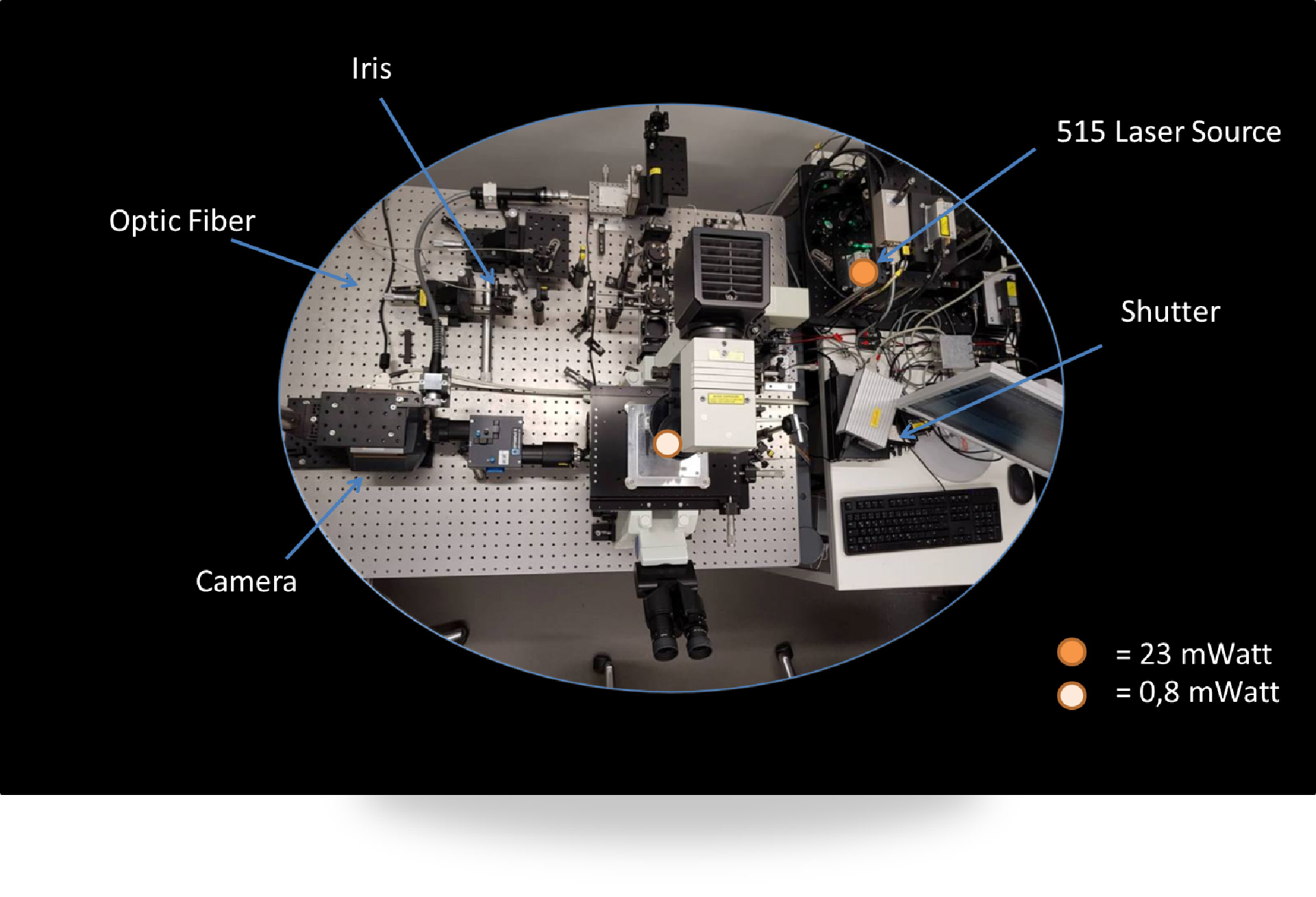
|
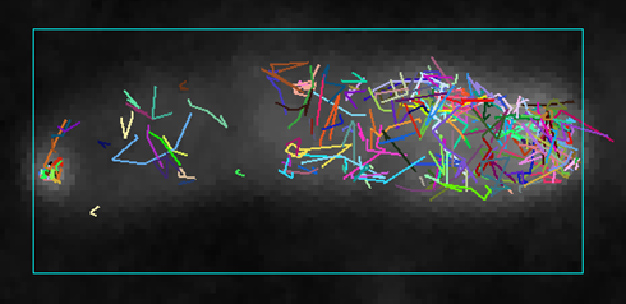

|
This work is also funded by the German Science Foundation (DFG; KO2184/8)
- Translational control of cellular stress response
The ribosome is a sophisticated cellular machine, composed of RNA and protein, which translates the mRNA-encoded genetic information into protein and thus acts at the centre of gene expression. Initially, the ribosome was considered to be an unchangeable entity, equipped with ribosomal components and translation factors for efficient and precise protein synthesis. However, this view is challenged by studies indicating that ribosome composition dynamically responds to intrinsic and extrinsic cues, which facilitates the cellular adaptation to a wide variety of environmental and metabolic conditions. We are in particular interested in understanding the role of the ribosome-associated protein YchF. YchF and its human homologue Ola1 have been implicated in multiple processes ranging from iron metabolism to tumorigenesis, but the molecular mechanism that allows YchF/Ola1 to control these diverse processes is unknown and currently under investigation.
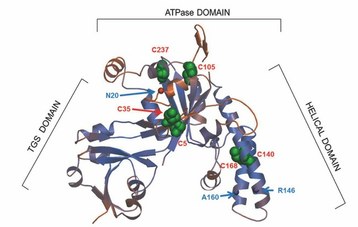 |
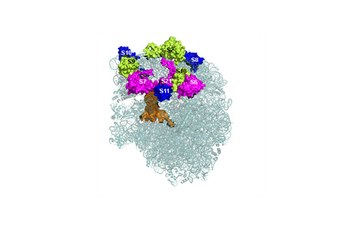 |
Structure of the universally conserved ATPase YchF (left) and its contact sites on the 30S ribosomal subunit (right); coloured ribosomal proteins have been found to be in contact with YchF
This work is funded by the DFG via the CRC/SFB 1381 (https://www.sfb1381.uni-freiburg.de/)
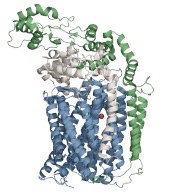
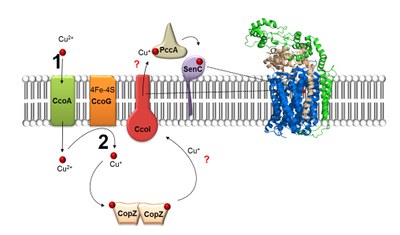
- Assembly of cytochrome cbb3-type cytochrome oxidases
Cytochrome cbb3-type cytochrome oxidases terminate the electron transfer chain in many pathogenic bacteria. Due to its high oxygen affinity, cbb3-type cytochrome oxidases are particularly important for bacteria that subsist in low oxygen environments, like the human gut. We are studying the assembly process of this protein complex, with a particular emphasis on the formation of the catalytic site of cbb3-type cytochrome oxidase, which contains two heme groups and one copper atom. The assembly of multi-subunit and several cofactor containing protein complexes is a highly complex process that requires sophisticated spatial and temporal coordination. cbb3-type cytochrome oxidase assembly has developed into a paradigm for such a process and is in particular interesting because it can identify novel targets for antibacterial therapy.
This work is funded by the DFG via the RTG2202 (https://www.taim.uni-freiburg.de/)
 |
 |
|
|||
Selected references:
Czech L, Mais CN, Sarmah P, Krazat H, Giammarinaro P, Freibert S, Musial J, Bernningshausen O., Steinchen W, Beckmann R, Koch HG, Bange, G. (2022) Shutdown of secretory pathway by the bacterial alarmones (p)ppGpp. Nature Commun. 13:1069. doi: 10.1038/s41467-022-28675-0.
Steinberg R, Origi A, Natriashvili A, Sarmah P, Licheva M, Walker PM, Kraft C, High S, Luirink J, Shi, WQ, Helmstädter M, Ulbrich MH, and Koch HG (2020) Post-translational insertion of small membrane proteins by the bacterial signal recognition particle. PloS Biology 18(9), e30000874, doi: 10.1371/journal.pbio.3000874.
Marckmann D, Trasnea PI, Schimpf J, Winterstein C, Andrei A, Schmollinger S, Blaby-Haas CE, Friedrich T, Daldal F, Koch HG (2019) The cbb3-type cytochrome oxidase assembly factor CcoG is a widely distributed cupric reductase. Proc. Natl. Acad. Sci. USA 116, 21166-21175 doi: 10.1073/pnas.1913803116.
Jauss B, Petriman NA, Drepper F, Franz L, Steinberg R, Warscheid B, Koch HG (2019) Non-competitive binding of PpiD and YidC to the SecYEG translocon expands the global view on the SecYEG interactome in E. coli. J. Biol. Chem. 294, 19167-19183 doi: 10.1074/jbc.RA119.010686.
Sachelaru, I., Winter, L., Knyazev, D., Zimmermann, M., Vogt, A., Kuttner, R., Ollinger, N., Siligan, C., Pohl, P., and Koch, H.G. (2017). YidC and SecYEG form a heterotetrameric protein translocation channel. Sci. Rep. 7, 101.
Denks, K., Sliwinski, N., Erichsen, V., Borodkina, B., Origi, A., and Koch, H.G. (2017). The signal recognition particle contacts uL23 and scans substrate translation inside the ribosomal tunnel. Nat. Microbiol. 2, 16215.
Braig, D., Nero, T., Koch, H.G., Kaiser, B., Wang, X., Thiele, J.R., Morton, C., Zeller, J., Kiefer, J., Potempa, L.A., Mellett, N.A., Miles, L.A., Du, X., Meikle, P.J., Huber-Lang, M., Stark, G.B., Parker, M.W., Peter, K., and Eisenhardt, S.U. (2017). Transitional changes in the CRP structure lead to the exposure of pro-inflammatory binding sites. Nat. Commun. 8, 14188.
Kuhn, P., Draycheva, A., Vogt, A., Petriman, N.A., Sturm, L., Drepper, F., Warscheid, B., Wintermeyer, W., and Koch, H.G. (2015). Ribosome binding induces repositioning of the signal recognition particle receptor on the translocon. J. Cell Biol. 211, 91-104.
Angelini, S., Deitermann, S., and Koch, H.G. (2005). FtsY, the bacterial SRP receptor functionally and physically interacts with the SecYEG translocon. EMBO Rep. 6, 476-481.
Alami, M., Lüke, I., Deitermann, S., Eisner, G., Koch, H.G., Brunner, J., and Müller, M. (2003). Differential interactions between a twin-arginine signal peptide and its translocase in Escherichia coli. Mol. Cell 12, 937-946.
Neumann-Haefelin, C., Schäfer, U., Müller, M., and Koch, H.G. (2000). SRP-dependent cotranslational targeting and SecA-dependent translocation analyzed as individual step in the export of a bacterial protein. EMBO J. 19, 6419-6426.

